Update on SARS-CoV-2 Variants
Update on SARS-CoV-2 Variants
Efficient Vaccine Distribution is Critical to Mitigate the Continuing Evolution of SARS-CoV-2
February 5, 2021
We’ve recently reported on SARS-CoV-2 variants, and described some of the reasons for concern. These include increased transmissibility, the possibility of false negative tests for viral RNA, resistance to immune protection from vaccine or prior infection, and pathogenicity. Rapid spread of variants makes it difficult to mitigate an ongoing pandemic.
UK Variant (B1.1.7)
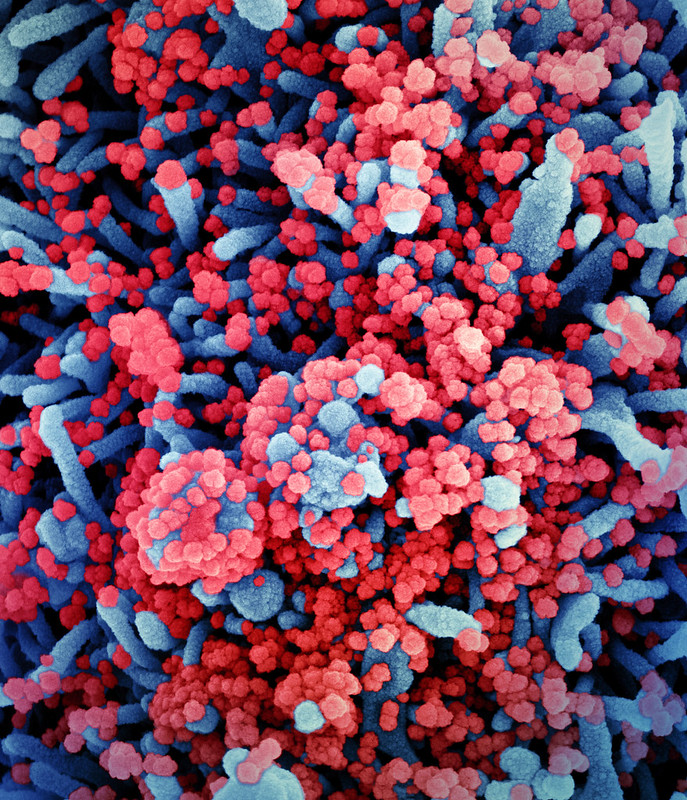
It appears that the B1.1.7 (UK lineage) variant can indeed cause false negatives on RT-PCR tests because of alterations in the gene for the spike protein, a phenomenon known as S-dropout(1). And it seems likely to be more transmissible(2), based at least in part upon the N501Y spike protein mutation conferring a greater affinity for the human ACE2 viral receptor as well as becoming the predominant strain in a short period of time. An N501Y substitution introduced into a wild type virus, however, does not alter the neutralization potency of vaccinee sera(3).
What about the other mutations in B1.1.7 and their possible effects on protective efficacy from vaccination or prior infection? This is unclear since the major types and levels of immunity and duration of required for protection are not precisely known. In addition, protective immunity is another important concern. Presumably, protection will involve some combination of innate immunity, neutralizing antibodies, and CD4+– and CD8+-T cell-mediated immunity. In fact, T cell-mediated immunity is orchestrating protective immunity of vaccines, including production of neutralizing antibody. Therefore, we need to consider not only changes in antibody epitopes but also changes in cytotoxic T-cell (CTL) epitopes and major histocompatibility (MHC) class I- and II-restricted epitopes. B1.1.7, has 19 mutations that change the amino acid composition of viral proteins, and many of these will affect antibody, CTL and MHC recognition. There are additional variants that arose independently from B1.1.7 in South Africa (called B.1.351) and Brazil (called P.1) that contain the same N501Y mutation in the spike protein, but are otherwise distinct from B1.1.7
Genomic Sequencing and Surveillance
Coronaviruses continually evolve, although they do so relatively slowly due to a viral proofreading enzyme. Thus, the more quickly and extensively SASR-CoV-2 spreads and the slower that universal vaccination is achieved, the greater the possibility of the appearance of vaccine resistant variants. Extensive genomic surveillance, transmission mitigation, and speed in vaccine distribution are critical.
The more quickly and extensively SASR-CoV-2 spreads and the slower that universal vaccination is achieved, the greater the possibility of the appearance of vaccine resistant variants.
If necessary, currently available vaccine platforms (i.e., mRNA-based and viral vectored vaccines) can also facilitate to update COVID-19 vaccines against emerging immune escape mutants.
Genomic sequencing analysis has identified some new and potentially troublesome variants. In this regard two recent preprints have reported naturally occurring mutations, with one report focused on possible effects on antibody binding(4) and the other on possible effects on CD8 CTL recognition(5). One study looked at available genomic sequences for mutations in the spike protein or near the binding interfaces between the spike protein receptor binding domain (RBD), the ACE2 receptor, and two human neutralizing antibodies derived from post-infection sera(4). In silico analyses suggested that 11 out of 16 of these naturally occurring mutations would disrupt binding by one of the two antibodies, but not to ACE2, while this was the case for 8 of the 12 mutations with the second antibody. Notably, all these mutations were observed in multiple independent natural isolates, suggesting they are likely to be representative of SARS-CoV-2 evolutionary strategies in humans.
Sequences containing MHC-I restricted CTL epitopes are similarly affected. Sequencing of 233 samples identified 197 non-synonymous changes in 27 CTL epitopes that are presented by two HLA subtypes, HLA-A*02:01 (a relatively common MHC subtype) and HLA-B*40:01(5). Identical mutations were often found in multiple samples. A survey of available SARS-CoV-2 sequences showed mutation prevalence in these epitopes ranging from ~0 to 7%. Serial samples showed that these variants tended to arise later in infection, suggesting that CTL activity was exerting evolutionary pressure on the virus, which implies that CTL activity is protective. Analyses of a subset of the mutant peptides showed reduced affinity for the HLA proteins mentioned above, suggesting that the mutations reflect an escape from CTL activity.
Other New Variants
Several other variants are increasingly spreading globally. One cluster that was first identified in Spain in June, 20A.EU1, is marked by 6 mutations, including A222V in the spike protein and A220V in the nucleoprotein(6). Yet another variant has emerged, notably in Santa Clara County, CA, that is marked by 3 signature mutations, including L452R in the spike protein. It is not clear whether these variants are indeed more infectious or more likely to escape vaccine-elicited immunity. In Denmark, variants containing a signature Y453F mutation in the spike protein passed between farmed minks and farm workers.
Conclusion
It is critical to extend our effort to genetic surveillance for monitoring the emergence of immune escape mutants against currently available COVID-19 vaccines. If necessary, currently available vaccine platforms (i.e., mRNA-based and viral vectored vaccines) can also facilitate to update COVID-19 vaccines against emerging immune escape mutants. No matter what, we need to keep in mind that vaccination can be the most efficient and effective measures in controlling the pandemics. Now, our public health strategy to fight the ongoing pandemic should be a combination of vaccination and continuation of wearing masks and maintaining physical distances.
- M. Kidd et al., S-variant SARS-CoV-2 is associated with significantly higher viral loads in samples tested by ThermoFisher TaqPath RT-QPCR. medRxiv, 2020.2012.2024.20248834 (2020).
- J. C. Santos, G. A. Passos, The high infectivity of SARS-CoV-2 B.1.1.7 is associated with increased interaction force between Spike-ACE2 caused by the viral N501Y mutation. bioRxiv, 2020.2012.2029.424708 (2021).
- X. Xie et al., Neutralization of N501Y mutant SARS-CoV-2 by BNT162b2 vaccine-elicited sera. bioRxiv, 2021.2001.2007.425740 (2021).
- E. Shang, P. H. Axelsen, The Potential for SARS-CoV-2 to Evade Both Natural and Vaccine-induced Immunity. bioRxiv, 2020.2012.2013.422567 (2020).
- B. Agerer et al., SARS-CoV-2 escapes CD8 T cell surveillance via mutations in MHC-I restricted epitopes. bioRxiv, 2020.2012.2018.423507 (2020).
- E. B. Hodcroft et al., Emergence and spread of a SARS-CoV-2 variant through Europe in the summer of 2020. medRxiv, 2020.2010.2025.20219063 (2020).
