Global Virus Network (GVN) Center of Excellence TSRI Investigation Reveals First-Ever Image of Elusive Viral Protein
Global Virus Network (GVN) Center of Excellence TSRI Investigation Reveals First-Ever Image of Elusive Viral Protein
New Direction for Lifesaving Vaccine
LA JOLLA, CA – June 2, 2017 – Before Ebola virus ever struck West Africa, locals were already on the lookout for a deadly pathogen: Lassa virus. With thousands dying from Lassa every year—and the potential for the virus to cause even larger outbreaks—researchers are committed to designing a vaccine to stop it.
Now a team led by Staff Scientist Kathryn Hastie and Professor Erica Ollmann Saphire at The Scripps Research Institute (TSRI) has solved the structure of the viral machinery that Lassa virus uses to enter human cells. Their study, published June 2, 2017 in the journal Science, is the first to show a key piece of the viral structure, called the surface glycoprotein, for any member of the deadly arenavirus family.
Importantly, the new structure provides a guide for designing a Lassa virus vaccine.
“Studying Lassa is critically important. Hundreds of thousands of people are infected with the virus every year, and it is the viral hemorrhagic fever that most frequently comes to the United States and Europe,” said Ollmann Saphire, senior author of the new study and co-director of the TSRI Global Virus Network (GVN) Center of Excellence. “This structure gives you the blueprints to make a vaccine that would deliver antibody protection.”
Like Ebola virus, Lassa fever starts with flu-like symptoms and can lead to debilitating vomiting, neurological problems and even hemorrhaging from the eyes, gums and nose. The disease is 50 to 70 percent fatal—and up to 90 percent fatal in pregnant women.
Success in Solving the Structure
The researchers solved this structure of the Lassa virus glycoprotein using a technique called x-ray crystallography, in which researchers prompt protein molecules from a virus to align and form a crystal. When x-rays hit the electrons in the crystal, they create a diffraction pattern that reveals the organization of the crystal and the molecular structure of the protein that formed it.
The researchers quickly found that Lassa virus posed a challenge for x-ray crystallography. The technique depends on having a stable protein, yet all the Lassa virus glycoprotein wanted to do was fall apart.
The problem was that glycoproteins are made up of smaller subunits. Other viruses have bonds that hold the subunits together, “like a staple,” Hastie said. Arenaviruses don’t have that staple; instead, the subunits float away from each other. Another challenge was to recreate part of the viral lifecycle in the lab—a stage when Lassa’s glycoprotein gets clipped into two subunits. “We had to figure out how to get the subunits to be sufficiently clipped and where to put the staple to make sure they stayed together,” Hastie said.
To overcome these obstacles, Hastie created mutant versions of important parts of the molecule and engineered a version of the Lassa virus surface glycoprotein that didn’t fall apart. She then used this model glycoprotein as a sort of magnet to find antibodies in patient samples that could bind with the glycoprotein to neutralize the virus.The molecular structure of the Lassa virus glycoprotein trimer provides the blueprints for vaccine design.
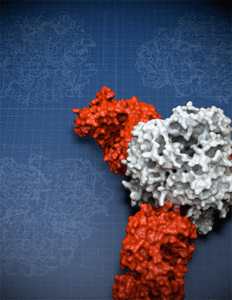
The molecular structure of the Lassa virus glycoprotein trimer provides the blueprints for vaccine design.
Image credit Christina Corbaci TSRI
At last, she solved the structure of the Lassa virus glycoprotein, bound to a neutralizing antibody from a human survivor.
Hastie’s structure showed that the glycoprotein has two parts. She compared the shape to an ice cream cone and a scoop of ice cream. A subunit called GP2 forms the cone, and the GP1 subunit sits on top. They work together when they encounter a host cell. GP1 binds to a host cell receptor, and GP2 starts the fusion process to enter that cell.
The new structure also showed a long structure hanging off the side of GP1—like a drip of melting ice cream running down the cone. This “drip” holds the two subunits together in their pre-fusion state.
Zooming in even closer, Hastie discovered that three of the GP1-GP2 pairs come together like a tripod. This arrangement appears to be unique to Lassa virus. Other viruses, such as influenza and HIV, also have three-part proteins (called trimers) at this site, but their subunits come together to form a pole, not a tripod.
“It was great to see exactly how Lassa was different from other viruses,” said Hastie, 10 years after starting the project. “It was a tremendous relief to finally have the structure.”
This tripod arrangement offers a path for vaccine design. The scientists found that 90 percent of the effective antibodies in Lassa patients targeted the spot where the three GP subunits came together. These antibodies locked the subunits together, preventing the virus from gearing up to enter a host cell.
A future vaccine would likely have the greatest chance of success if it could trigger the body to produce antibodies to target the same site.
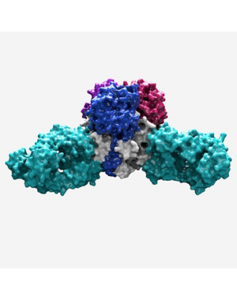
Antibody from a human survivor (turquoise) is shown inactivating the Lassa virus surface protein. The work shows how to engineer vaccine strategies to elicit protective immune responses. Image credit Christina Corbaci TSRI
Moving Forward with a Lassa Vaccine
The next step is to test a vaccine that will prompt the immune system to target Lassa’s glycoprotein. As director of the Viral Hemorrhagic Fever Immunotherapeutic Consortium, Ollmann Saphire is already coordinating with her partners at Tulane and Kenema to bring a vaccine to patients.
The Coalition for Epidemic Preparedness Innovations (CEPI), an international collaboration that includes the Wellcome Trust and the World Health Organization as partners has recently named a vaccine for Lassa virus as one of its three top priorities. “The community is keenly interested in making a Lassa vaccine, and we think we have the best template to do that,” said Ollmann Saphire.
She added that with Hastie’s techniques for solving arenavirus structures, researchers can now get a closer look at other hemorrhagic fever viruses, which cause death, neurological diseases and even birth defects around the world.
In addition to Ollmann Saphire and Hastie, authors of the study, “Structural basis for antibody-mediated neutralization of Lassa virus,” were Michelle A. Zandonatti of TSRI; James E. Robinson and Robert F. Garry of Tulane University and Director of Tulane’s GVN Center of Excellence; Lara M. Kleinfelter and Kartik Chandran of the Albert Einstein College of Medicine; and Megan L. Heinrich, Megan M. Rowland and Luis M. Branco of Zalgen Labs.
The study was supported by the National Institutes of Health (grants 1U19AI109762-01, R21 AI116112 and contract HHSC272200900049C) and an Investigators in Pathogenesis of Infectious Diseases Award from the Burroughs Wellcome Fund.
TSRI and Tulane University are Centers of Excellence in the Global Virus Network (GVN), which represents 38 Centers of Excellence and six affiliates in 24 countries and comprises foremost experts in every class of virus causing disease in humans. The GVN is a global authority and resource for the identification and investigation, interpretation and explanation, control and suppression, of viral diseases posing threats to mankind. It enhances the international capacity for reactive, proactive and interactive activities that address mankind-threatening viruses and addresses a global need for coordinated virology training through scholarly exchange programs for recruiting and training young scientists in medical virology. The GVN also serves as a resource to governments and international organizations seeking advice about viral disease threats, prevention or response strategies, and GVN advocates for research and training on virus infections and their many disease manifestations.
