
Diagnostic Efficacy
- The current molecular tests detect most of the variants and thus are able to diagnose COVID-19 infection by such variants. Yet, the fine identification of the type of variants is still based on sequence analysis although multiplex PCR test are being evaluated.
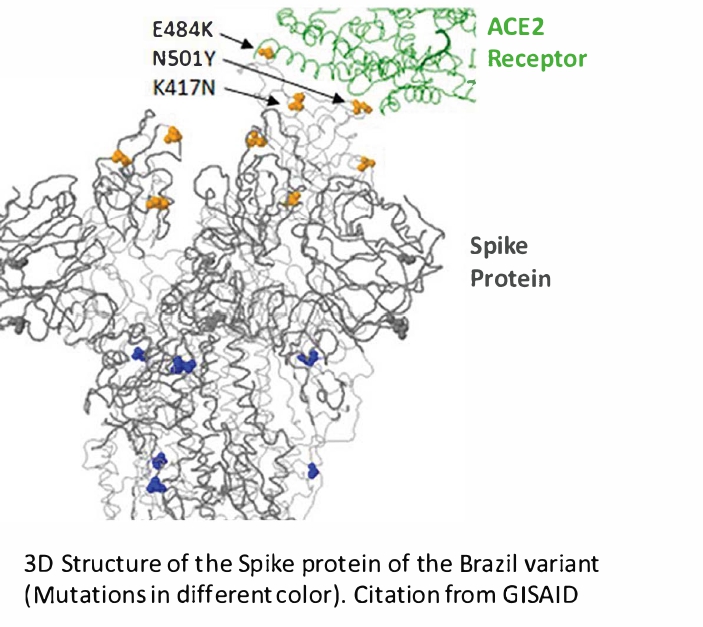
- Indeed, the current variants of concern show distinctive mutations in the spike protein. Due to such mutations, most diagnostic tests for COVID-19 have been designed by targeting not only the spike protein but also other conserved proteins. For example, molecular tests designed to detect multiple SARS-CoV-2 genes (i.e., multiplex reverse transcription polymerase chain reaction targeting ORF1ab, N, and E genes) are less susceptible to the effects of genetic variation than tests designed to detect a single gene. The FDA is also monitoring the potential effects of genetic variation in molecular tests that have received Emergency Use Authorization and provides information about the tests (https://www.fda.gov/medical-devices/letters-health-care-providers/genetic-variants-sars-cov-2-may-lead-false-negative-results-molecular-tests-detection-sars-cov-2).
- Overall, the precise characterization of the variants still relies on genomic sequencing analysis. For instance, CDC is currently increasing sequence surveillance to more than 6000 samples per week to efficiently monitor the variants of concerns and other emerging variants. COVID-19 caused by variants in the U.S. can be found here.
